Patterns in Permutations
In this column, we will consider classes of permutations that appear in various contexts and seek a means of describing the permutations in these classes. ...
Introduction
Permutations appear so commonly in mathematics that even elementary students are familiar with them. In this column, we will look at some classes of permutations and see how they may be described using the language of patterns in permutations.
Of course, a permutation is usually thought of as a bijection $\sigma$ from a set having $n$ elements to itself. Frequently, we think of $\sigma:\{1,2,\ldots,n\}\to\{1,2,\ldots,n\}$ and write $\sigma = \sigma_1\sigma_2\ldots\sigma_n$ to mean $\sigma(i) = \sigma_i$. For example, the permutation $\sigma = 52314$ has the property that $$ \sigma(1) = 5, \sigma(2) = 2, \sigma(3) = 3, \sigma(4) = 1,\sigma(5) = 4. $$
In this column, we will consider classes of permutations that appear in various contexts and seek a means of describing the permutations in these classes. Let's begin with a classical problem introduced by Knuth before considering a more recent problem in computational biology.
Sorting with a stack
We begin by looking at how a stack, a simple but powerful data structure frequently used in computing, may be used for sorting. For example, we will input a permutation of the integers $1,2,3,\ldots, n$, such as 3124, and seek a process that sorts the integers into the increasing sequence 1234.
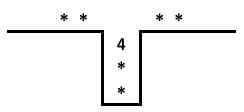
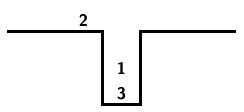
A stack may be schematically represented as shown below.
At the beginning of the sorting process, we have the input on the right.
Individual numbers are transferred from the input to the output with the stack acting as a way station. There are only two allowed operations.
- Numbers may be pushed from the left of the input string to the top of the stack.
- Numbers may be popped from the top of the stack to the right of the output string.
The following rules define an algorithm that attempts to sort the input into a string of increasing integers.
- The algorithm ends when the stack and the input are empty.
- Perform a push if the stack is empty and input is not empty.
- Perform a pop if the input is empty and the stack is not empty.
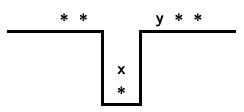
- If $x$ is on top of the stack and $y$ on the left of the input,
perform a pop if $x \lt y$
and a push if $x \gt y$.
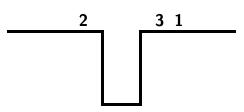
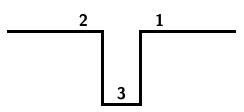
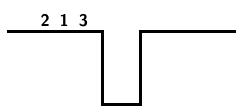
The following steps result from applying this algorithm to the input string 3124:
Comments?
Stack-sortable permutations
This example shows that our stack-sorting algorithm is capable of sorting the permutation 3124. Can every permutation be sorted in this way? An example with input 231 shows that the answer is no.
Since our algorithm does not sort the permutation 231 into 123, there are clearly some permutations that can be sorted in this way and some that can't. A permutation is called stack-sortable if it can be sorted using a stack. In searching for a criterion that detects when a permutation is stack-sortable, we are led to the notion of patterns in permutations.
First, we will introduce some terminology. A subsequence of a permutation $\sigma$ is a list of numbers obtained by removing some of the numbers from $\sigma$. For instance, the permutation $563412$ contains the subsequence $5342$ by removing $1$ and $6$.
The reduced form of a subsequence $i_1i_2\ldots i_p$ is the permutation $\pi$ obtained by relabeling the elements in $\{i_1, i_2,\ldots, i_p\}$ with the elements of $\{1,2,\ldots, p\}$ in an order-preserving way. For instance, the subsequence $4532$ has the reduced form $3421$.
We say that a permutation $\sigma$ contains the pattern $\pi$ if $\pi$ is the reduced form of a subsequence of $\sigma$. For instance, the permutation $563412$ contains the pattern $312$, which appears as the reduced form of the subsequences $534$, $512$, $634$, $612$, $312$, and $412$.
We denote the fact that $\pi$ is a pattern of $\sigma$ by writing $\pi\preceq\sigma$. If, in addition, $\pi$ is not equal to $\sigma$, we write $\pi\prec\sigma$.
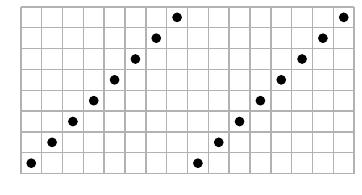
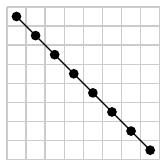
It is sometimes helpful to represent a permutation $\sigma$ as the graph of a function $\{1,2,\ldots,n\}\to\{1,2,\ldots,n\}$. For instance, the permutation $\sigma=563412$ may be represented as
It is now visually clear that the pattern 312 is realized by the subsequence 534.
Let's now return to our discussion about which permutations can be sorted with a stack. We have seen that the permutation 231 cannot be sorted. Now we can say that any permutation containing the pattern 231 cannot be sorted.
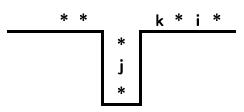
For instance, suppose $\sigma$ is a permutation containing the pattern 231. This means that we have $i\lt j\lt k$ appearing in the permutation as
Of course, there may be more than one appearance of the 231 pattern so let's choose one such that the distance between the appearances of j and k is as small as possible. Looking at the graph of the permutation, this means there are no elements in the red shaded region.
By the time that k appears on the right of the input string, j has already gone onto the stack. If it is still there, then any elements above j are smaller than k. Therefore, we will perform a sequence of pop operations until j is popped.
This means that j will appear in the output string before i, even though i $\lt$ j, and so the output string is not sorted. This shows that a permutation containing the 231 pattern is not sortable.
In fact, we can say more. Suppose that a permuation is not sortable, meaning there is a pair i $\lt$ j such that j appears in the output string before i. This means that j is popped off the stack before i by some element k, which appears between j and i in the input.
It must be true that j $\lt$ k so that we have i $\lt$ j $\lt$ k. We have now found an appearance of the 231 pattern.
This shows that
A permutation is stack-sortable if and only if it contains no appearance of the 231 pattern.
Comments?
How many stack-sortable permutations are there?
Now that we have a criterion that tells us which permutations are stack-sortable, it's easy to count the number of these permutations.
We will denote the number of stack-sortable permutations of length $n$ by $s_n$. It is easy to see that $s_1 = 1$, $s_2 = 2$, and $s_3 = 5$. We will also adopt the convention that $s_0 = 1$.
Let's suppose that $\sigma$ is a permutation of length $n$ with $\sigma_i = n$. As illustrated below, if $\sigma$ is to avoid the 231 pattern, then $\sigma_1\sigma_2\ldots\sigma_{i-1}$ must be a permutation of $\{1,2,\ldots, i-1\}$ avoiding the 231 pattern and $\sigma_{i+1}\sigma_{i+2}\ldots\sigma_n$ is a permutation of $\{i,i+1,\ldots,n-1\}$ avoiding the 231 pattern.
This shows that $$ s_n = \sum_{i=0}^{n-1} s_is_{n-1-i}, $$ which is the well-known recurrence relation that defines the Catalan numbers $C_n$, and therefore $$ s_n = C_n = \frac{1}{n+1}{2n\choose{n}}. $$
Pattern avoidance
So far, we have been looking at the set of stack-sortable permutations and have found they are characterized by the absence of the pattern 231. As this is a typical phenomenon, we will introduce some terminology to help our discussions.
First, suppose that $\tau\preceq\pi$ and $\pi\preceq\sigma$. A little thought shows that $\tau\preceq\sigma$ as well. In other words, $\preceq$ is a transitive relation.
Suppose now that $\sigma$ is a stack-sortable permutation so that 231 is not a pattern in $\sigma$. It therefore follows that any pattern $\pi\preceq\sigma$ is also stack-sortable. In this case, we say that the stack-sortable permutations form a permutation class. More generally, a permutation class $\cal C$ is a set of permutations such that if $\sigma\in{\cal C}$ and $\pi\preceq\sigma$, then $\pi\in{\cal C}$ as well.
We say that ${\cal B}$ is a basis for a permutation class ${\cal C}$ if $\pi\in{\cal B}$ is not in ${\cal C}$ but any pattern $\sigma\prec\pi$ is in ${\cal C}$. In other words, a basis for a permutation class consists of a minimal set of permutations not in that class. For instance, we have seen that a basis for the class of stack-sortable permutations consists of the single permutation 231.
A problem in computational biology
Now that complete genome sequences are readily available, it is possible to compare two genomes and ask how far apart they are under various rearrangement strategies. Evolutionary biologists have identified a particular model of genome rearrangement, known as tandem duplication-random loss, that explains many observed rearrangements in mitochondrial genomes. As we will see following Chaudhuri et al and Bouvel and Pergola, this model of rearrangement leads to a permutation class whose basis may be easily identified.
A rearrangement of a genome may be modeled by a permutation. We begin by labeling the genes in a sequence by 1, 2, 3, $\ldots$, n.
In the first phase of the rearrangment, this sequence of genes is duplicated to obtain a sequence twice as long as the original.
Notice that each gene now appears twice. In the second phase of the rearrangement, one copy of each gene is randomly selected for removal,
which leads to the final rearrangement.
We are interested in studying the permutations that result after applying a certain number $p$ of duplication-loss rearrangements, as $p$ may be thought of as measuring the evolutionary distance between the two genomes. To this end, we introduce ${\cal C}_p$, the class of permutations obtained by $p$ duplication-loss rearrangements and ask to find a basis for it.
The key idea is to focus on the number of descents in the permutations. In the permutation $\sigma$, we say that there is a descent at position $i$ if $\sigma_i \gt \sigma_{i+1}$. (If $\sigma_i\lt\sigma_{i+1}$, then we say there is an ascent at position $i$.) Notice that the permutation we found from one duplication-loss rearrangement, shown above, has one descent.
Suppose now that we perform a second duplication-loss rearrangement. After duplicating, we have three descents: the original descent is duplicated, and a third is introduced.
After removing one randomly selected copy of each gene, we arrive at the final permutation, which again has three descents.
Of course, it is possible that a descent is removed in the loss phase of the rearrangement, but we see that there can be no more than three descents.
In the same way, we see that, after $p$ duplication-loss rearrangements, there may be no more than $d=2^p-1$ descents. In fact, Chaudhuri et al showed that this property characterizes ${\cal C}_p$.
${\cal C}_p$ is the class of permutations with at most $2^p-1$ descents.
A basis for ${\cal C}_p$ is then ${\cal B}_d$, the permutations with $d=2^p$ descents that are minimal in this sense: if $\sigma\in{\cal B}_d$ and $\pi\prec\sigma$, then $\pi$ has fewer than $d$ descents.
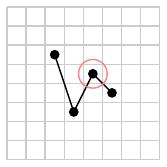
It is straightforward to see that the basis ${\cal B}_d$ is finite: as seen below, a permutation having two adjacent ascents cannot be minimal since removing the circled element produces a subsequence having the same number of descents.
In the same way, a minimal permutation cannot begin or end with an ascent. Therefore, the number of ascents $a$ must be less than the number of descents $d$; that is, $a\leq d-1$.
If the length of the permutation is $n$, then $a + d = n-1$, which means that a minimal permutation has $n\leq 2d$. The shortest permutation having $d$ descents is an inversion of length $d+1$ while alternating permutations have the longest possible length $2d$.
Therefore, the length of a permutation in ${\cal B}_d$ is bounded, which implies that ${\cal B}_d$ is finite.
We now define a more refined type of pattern, known as a vincular pattern. We say that $\pi\prec\sigma$ is a vincular pattern if the subsequence in $\sigma$ that forms $\pi$ consists of adjacent numbers. This is denoted by underlining the numbers in $\pi$.
For instance, in the permutation $\sigma = 53421$, the subsequence 342 is an appearance of the vincular pattern $\underline{231}$ since the numbers are adjacent. The numbers 341 form an appearance of 231 in $\sigma$, but not $\underline{231}$. Descents are appearances of the vincular pattern $\underline{21}$ while ascents are $\underline{12}$.
Since the permutations in ${\cal B}_d$ are minimal, the following vincular patterns cannot appear since the circled number may be removed without lowering the number of descents.
In fact, the only forms in which ascents may occur are as shown below:
Therefore, ascents in permutations in ${\cal B}_d$ manifest as appearances of vincular patterns $\underline{2143}$ and $\underline{3142}$.
This gives the following characterization of permutations in the basis ${\cal B}_d$:
A permutation is in ${\cal B}_d$ if and only if it has exactly $d$ descents and any ascents appear in the patterns $\underline{2143}$ and $\underline{3142}$.
The Catalan numbers make another appearance: Bouvel and Pergola show that the number of minimal permutations having $d$ descents and length $2d$ is the Catalan number $C_d$.
Comments?
Summary
The use of patterns to describe permutation classes is currently a growing area. In "A Singular Mathematical Promenade," Ghys describes a recent result of Kontsevich concerning "polynomial interchanges."
Suppose we have a collection of polynomials ordered so that $P_1(x) \lt P_2(x) \lt \ldots \lt P_n(x)$ for negative values of $x$ close to 0. We obtain a permutation by comparing this ordering to that obtained using positive values of $x$ close to 0.
For instance, the polynomials shown below define the permutation 213.
Kontsevich showed that the class of permutations that arise in this way avoid the patterns 2413 and 3142.
Kitaev's book serves to systematize vocabulary and notation in addition to demonstrating a wealth of areas in which these ideas appear from theoretical computer science and discrete mathematics to algebraic geometry and theoretical physics.
Comments?
References
-
Donald E. Knuth. The art of computer programming. Volume 1, Fundamental Algorithms. Addison-Wesley, Reading, 1975.
-
Sergey Kitaev. Patterns in Permutations and Words. Monographs in Theoretical Computer Science. Springer, Heidelberg, 2011.
-
Étienne Ghys. A singular mathematical promenade. Freely available through the American Mathematical Society's Open Math Books.
As the title suggests, Ghys's book is enjoyable tour through a remarkable range of mathematical topics.
-
Bridget Tenner. Database of Permutation Pattern Avoidance.
Similar to the Online Encyclopedia of Integer Sequences, this database allows one to enter a set of patterns and find phenomena leading to permutation classes that avoid those patterns.
-
Mathilde Bouvel and Elisa Pergola. Post and permutations in the duplication-loss model: Minimal permutations with $d$ descents. Theoretical Computer Science. Volume 411, 2487-2501, 2010.
-
Kamalika Chaudhuri, Kevin Chen, Radu Mihaescu, and Satish Rao. On the tandem duplication-random loss model of genome rearrangement. SODA, pages 564–570, 2006.


 David Austin
David Austin